
Help
About trRosettaRNA
[Input Preparation] Shown in Figure 1, the input to trRosettaRNA is the nucleotide sequence of the target RNA. Optionally, users can also input their custom multiple sequence alignment (MSA) and/or secondary structure (SS).
[Structure Prediction] The generated/input MSA and SS are fed into an end-to-end neural network to predict initial 3D atomic coordinates and 2D internucleotide distances.
[Structure Optimization] Subsequently, the predicted coordinates are then optimized through fast relaxation or energy minimization (min.) (depending on the severity of steric clash) to produce a physically realistic structure.
[Final Outputs] The final outputs include the predicted 3D structure and an associated confidence score (pLDDT).
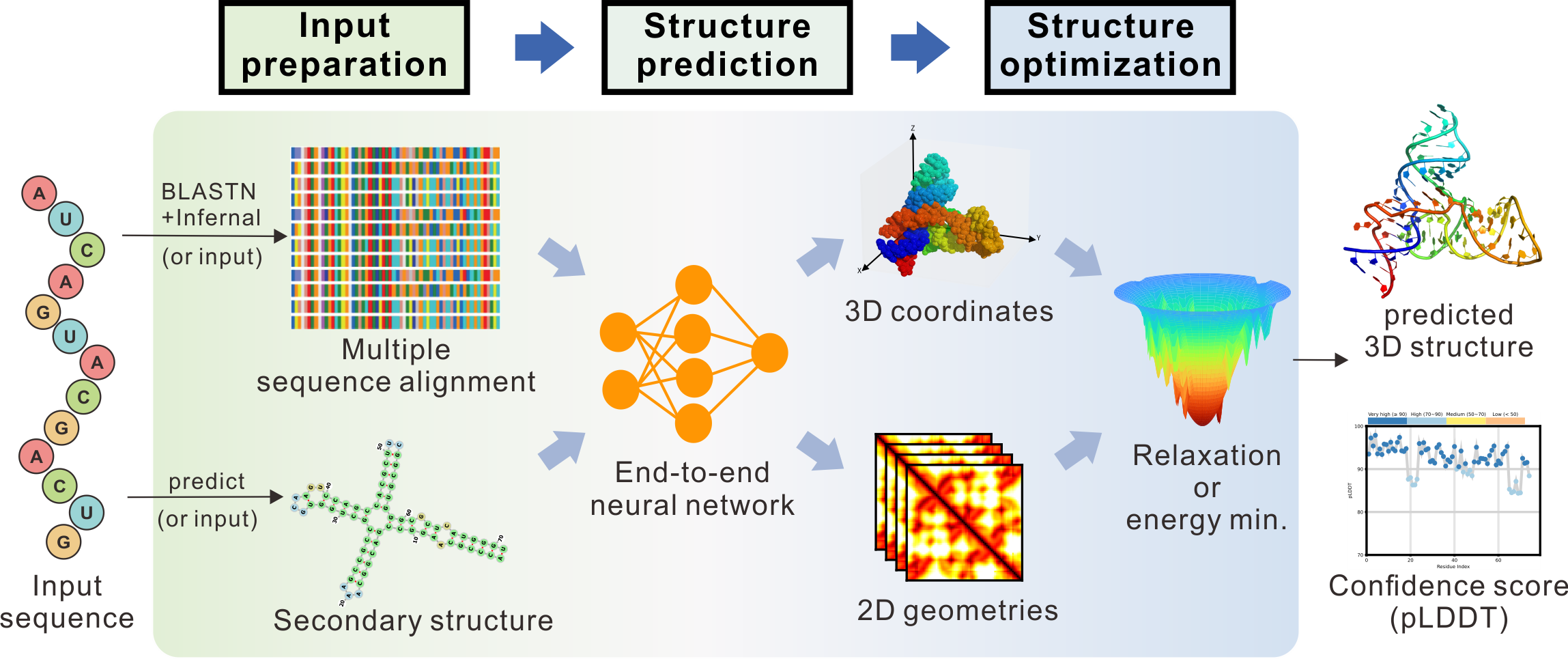
Figure 1. The flowchart of the trRosettaRNA server.
trRosettaRNA has been validated in blind tests, including CASP and RNA-Puzzles (Figure 2). These results suggest that its automated predictions achieve state-of-the-art performance among automated approaches, and are highly competitive with those of top human groups on natural RNAs.

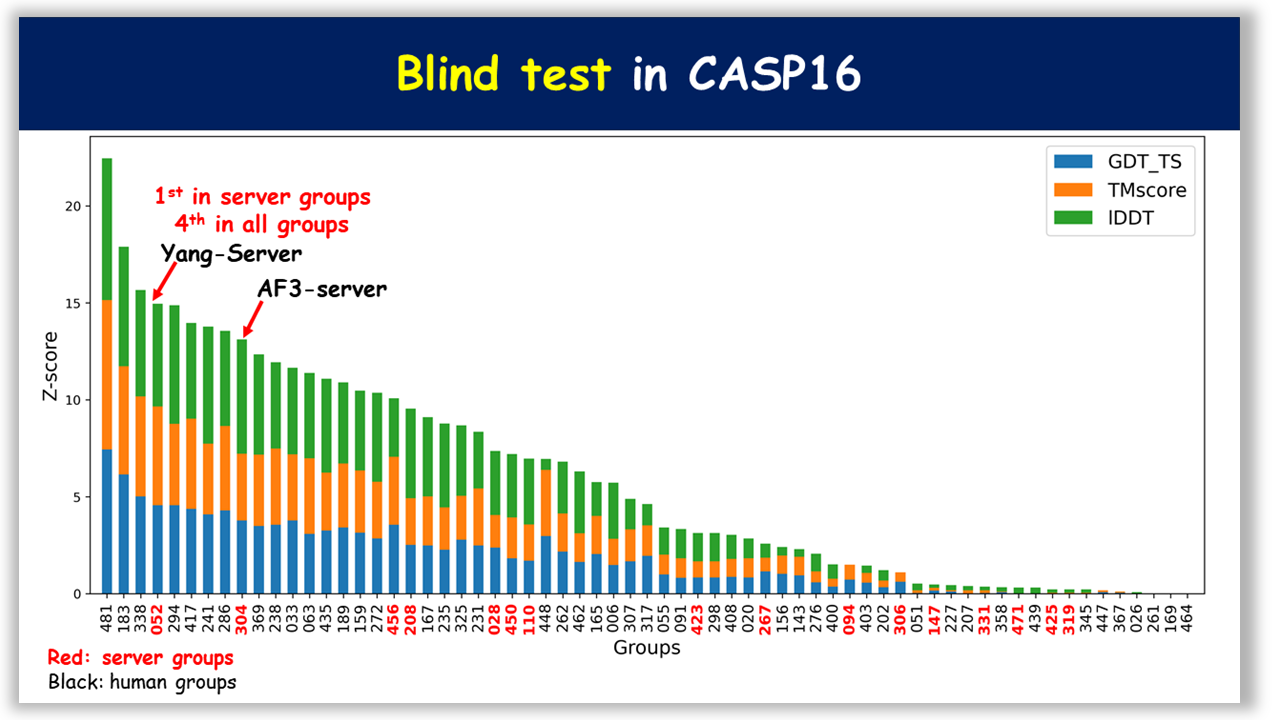
Figure 2. Blind test results in CASP and RNA-Puzzles.
The trRosettaRNA server runs BLASTN and Infernal on the RNAcentral database to generate MSA.
At the end of trRosettaRNA neural network, we use a simple ResNet module to predict the lDDT(Local Distance Difference Test) of the predicted structure model. The lDDT score ranges from 0 to 100, with higher values indicating better accuracy.
Figure 3 shows the relationship between the predicted (pLDDT) and the real lDDT for modeling on 50 RNAs used for benchmark tests. The Pearson correlation coefficients for global and local metrics are 0.798 and 0.736, respectively.
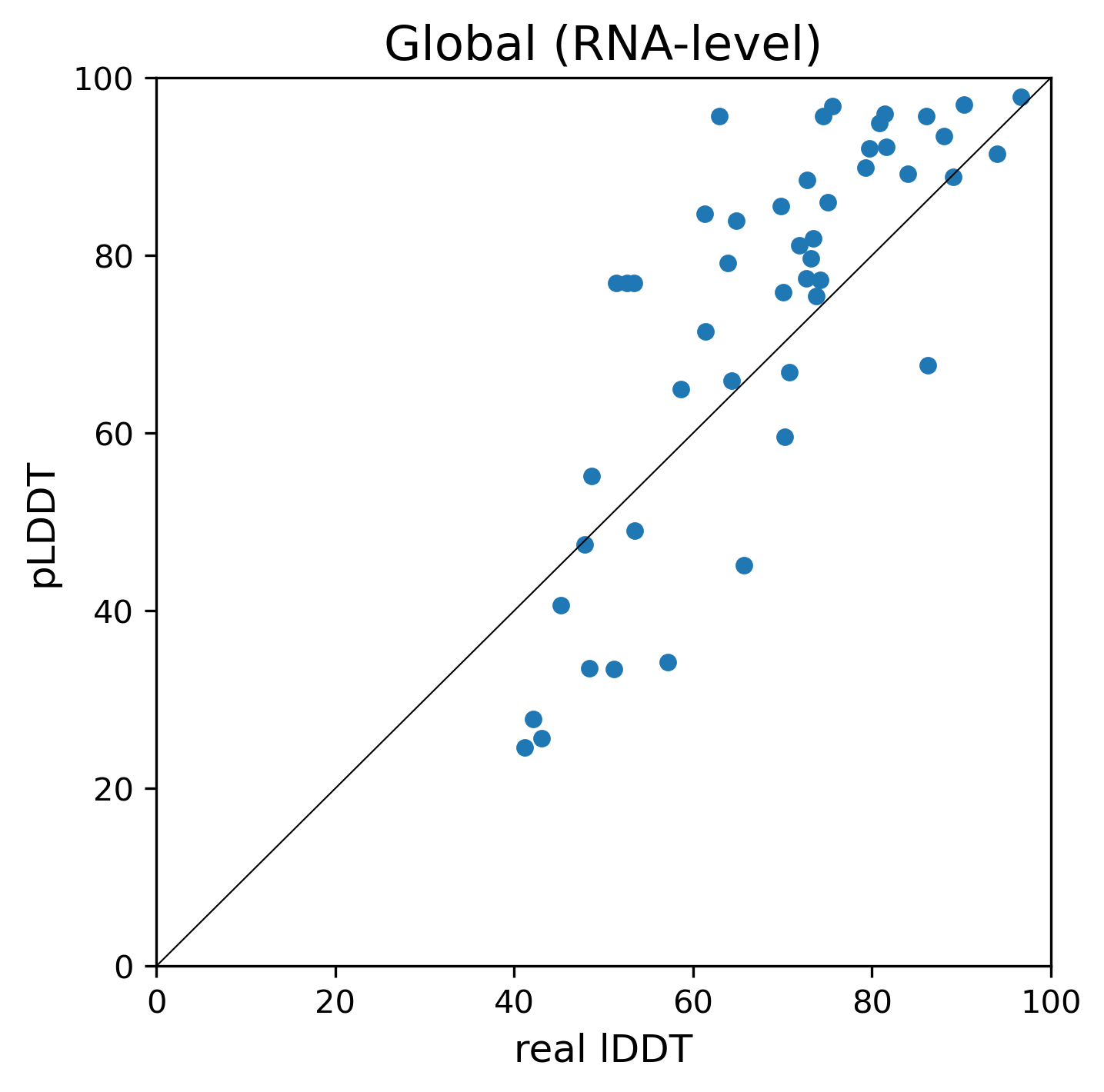
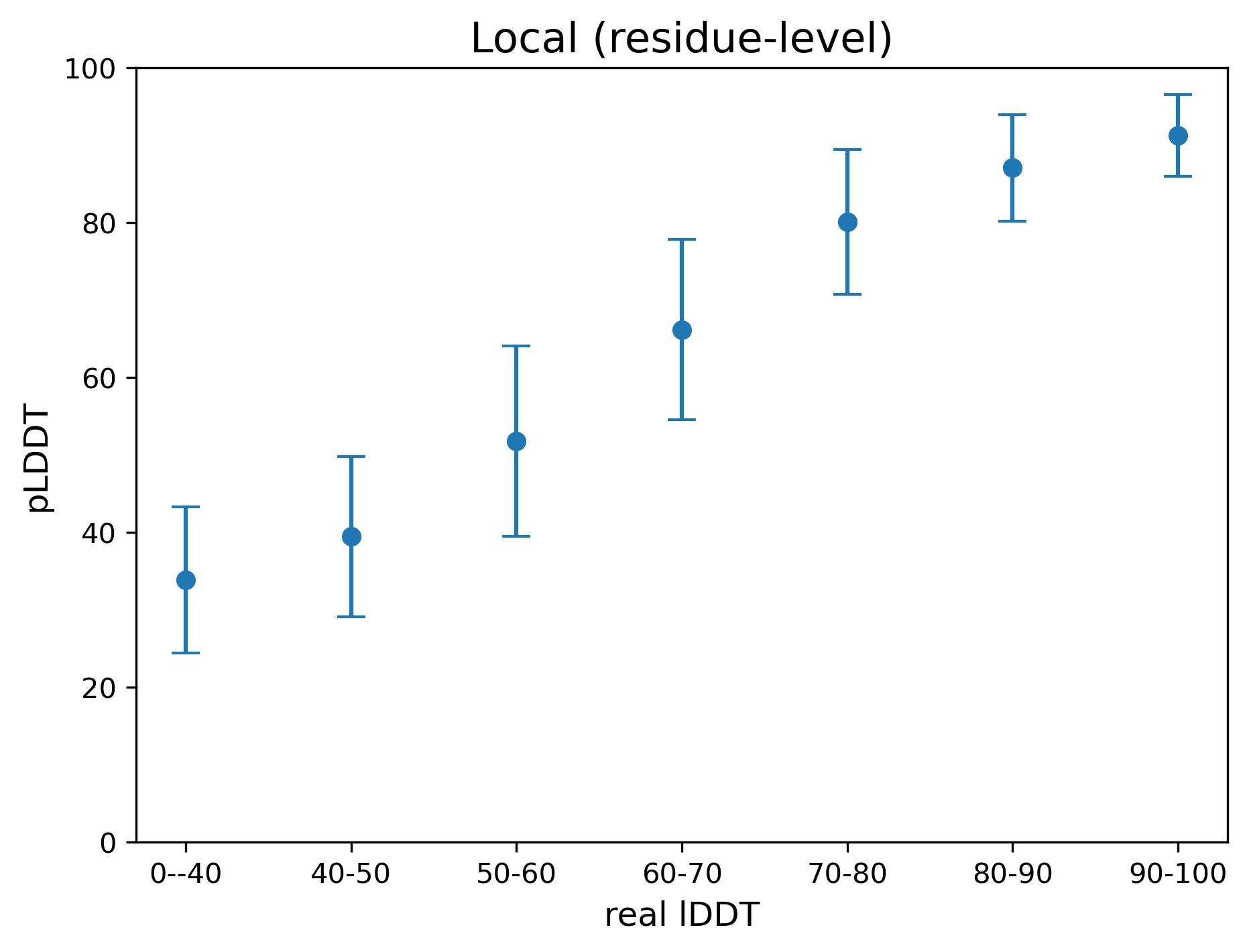
Figure 3. The relationship between the predicted lDDT (pLDDT) and the real lDDT.
Job submission
Job submission guide:(1) Input a RNA sequence in FASTA format or a MSA in A3M/FASTA/A2M/STO format.
(2) Specify your input type.
(3) (Optional) Provide your custom secondary structure in dot-bracket/CT format (If not provided, the secondary structure will be predicted by trRNA-SS).
(4) (Optional) Provide your email address.
(5) (Optional) Assign your target name.
(6) Choose whether to keep your results private.
(7) Submit.
Notice: Due to computing resource limitation, we now allow no more than 20 running/pending jobs per user at the same time.
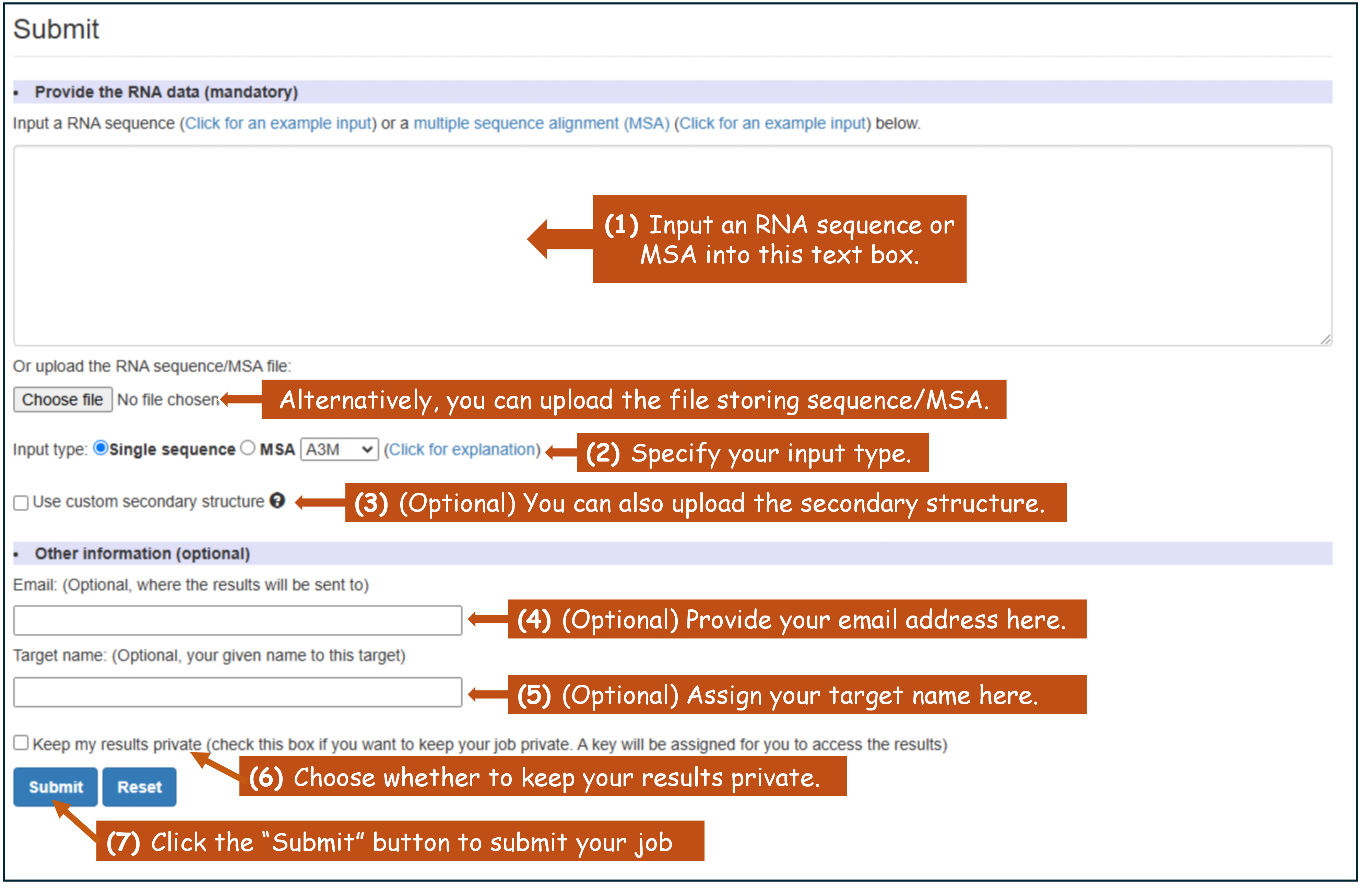
Figure 4. The "Submit" section in trRosettaRNA home page.
Output explanation
The trRosettaRNA modeling results are generally summarized in a webpage, the link of which is sent to the user upon job completion if the email address has been provided during submission(see an example of the trRosettaRNA output). A tallbar file containing the key modeling results can be downloaded from the top of this page.This section contains:
(1) Predicted 3D model visualization.
(2) Model quality estimation.
(3) Modeling method description.
(4) Separate download links for predicted structure, MSA (multiple sequence alignment) and inter-nucleotide distances.
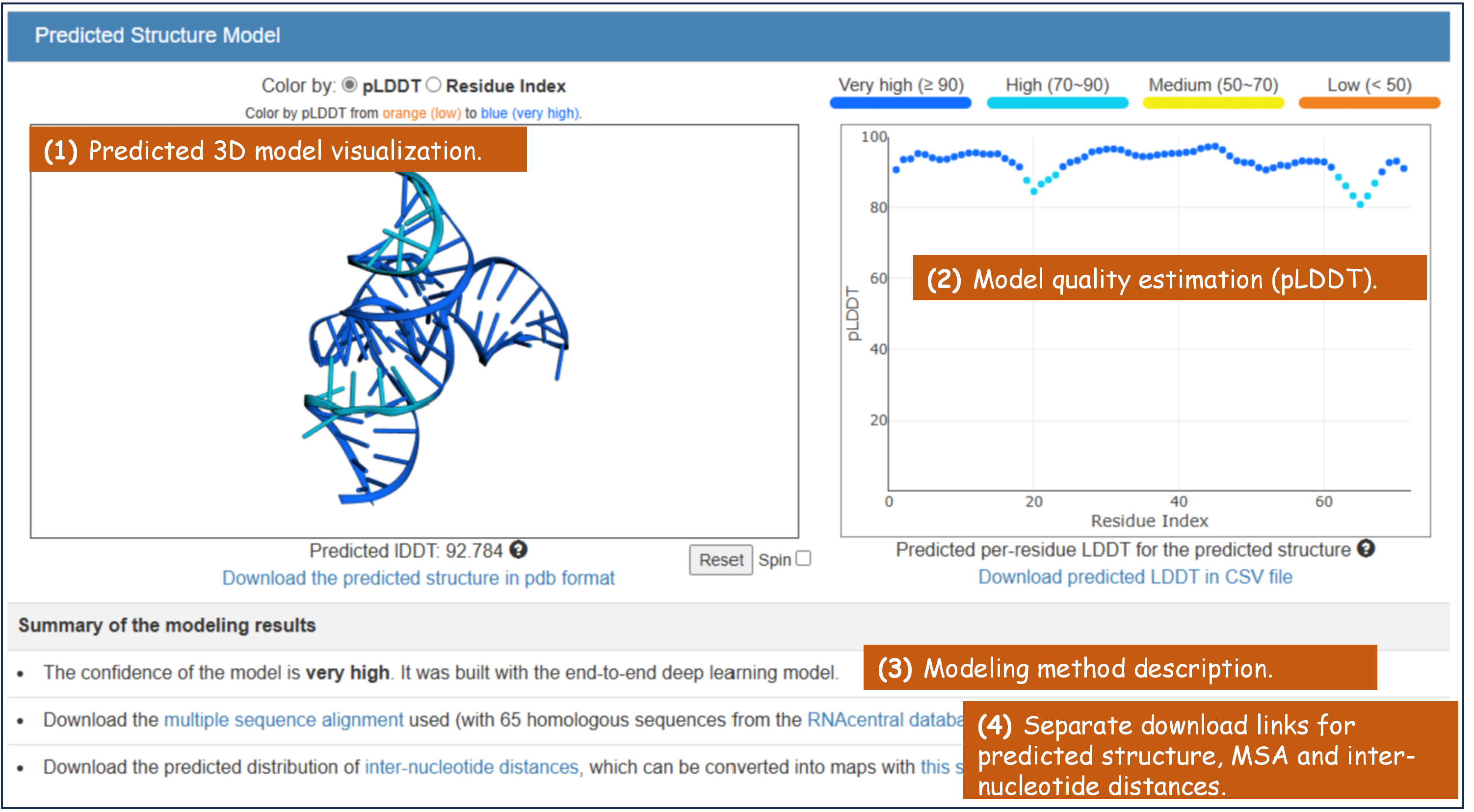
Figure 5. The "Predicted Structure Models" section in trRosettaRNA result page.
(1) Contact map, which displays the predicted probability of nucleotide pairs being in contact, i.e., the minimum distance between all atoms of them is less than 8Å.
(2) Distance map, which displays the predicted distance (3 - 40 Å) between nucleotide pairs.
Figure 6. The "Predicted 2D Information" section in trRosettaRNA result page.
(1) predicted by trRNA-SS or input by user.
(2) extracted from the predicted 3D structure.
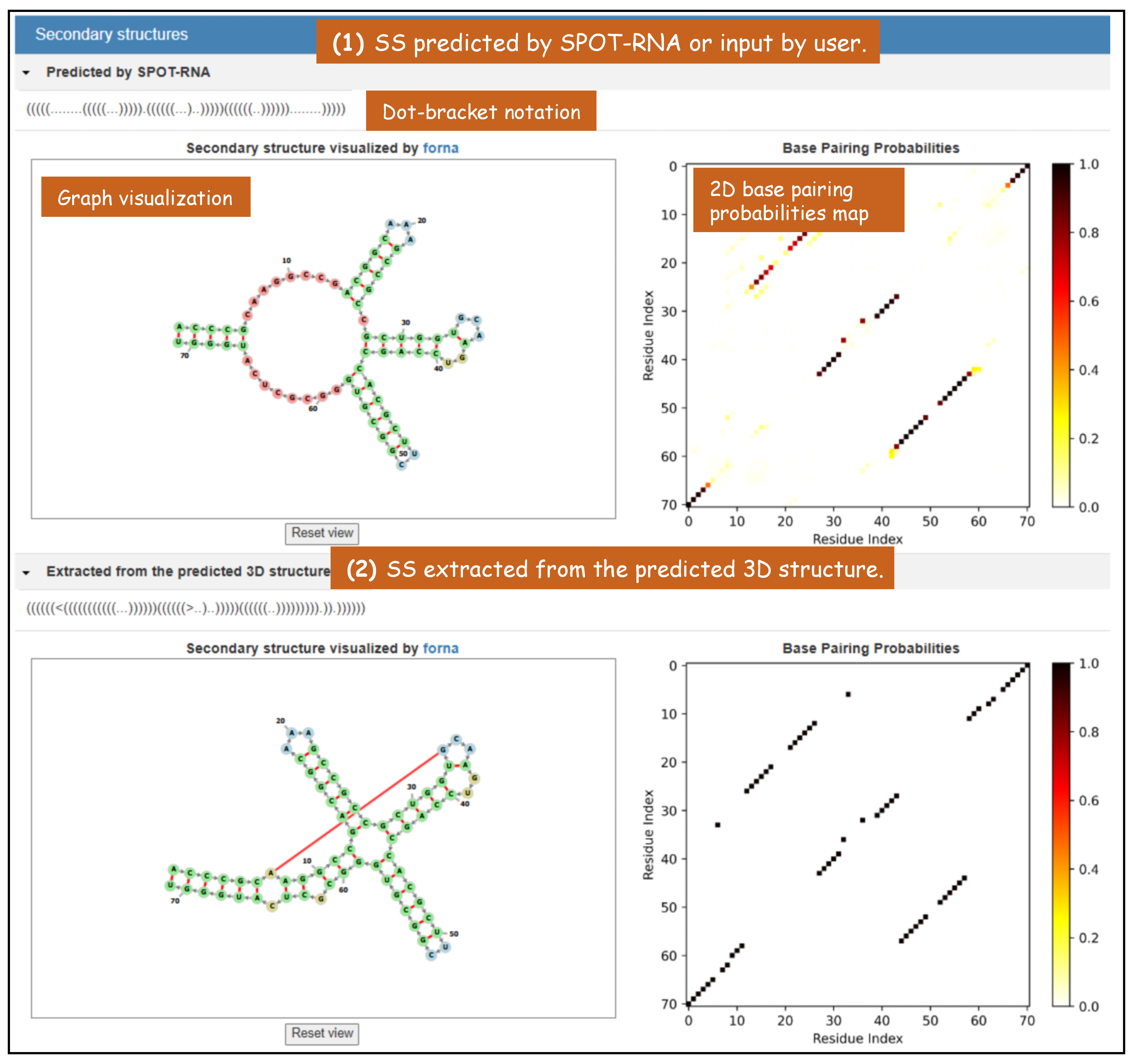
Figure 7. The "Secondary structures" section in trRosettaRNA result page.